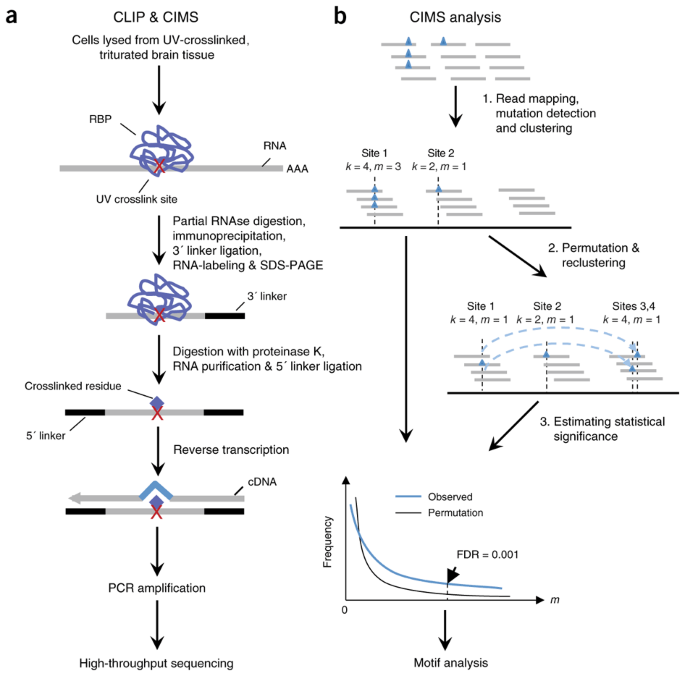
Mapping in vivo protein-RNA interactions at single-nucleotide resolution from HITS-CLIP data | Nature Biotechnology
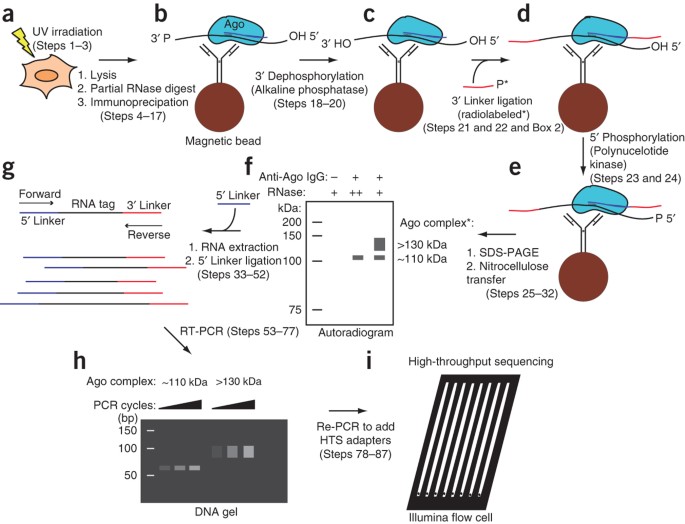
Mapping Argonaute and conventional RNA-binding protein interactions with RNA at single-nucleotide resolution using HITS-CLIP and CIMS analysis | Nature Protocols
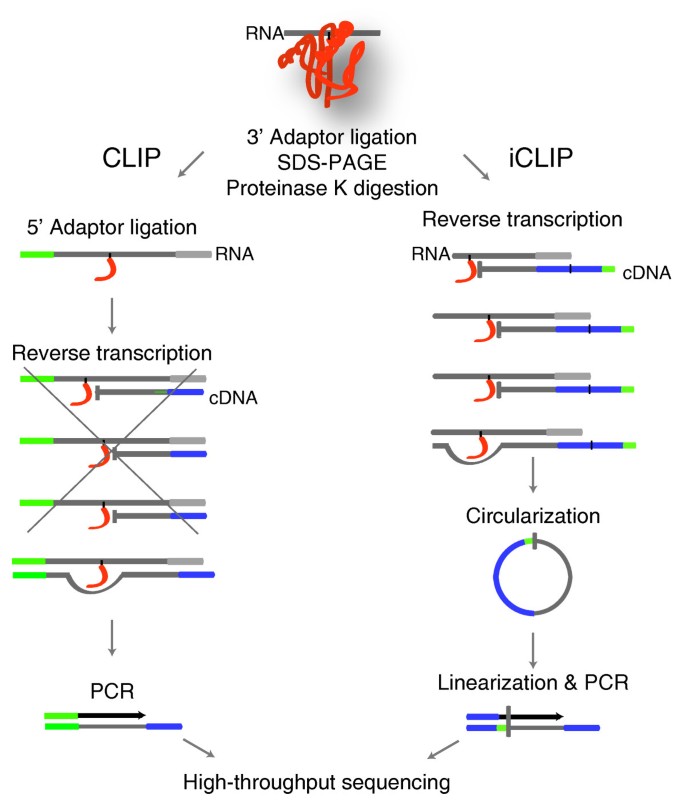
Analysis of CLIP and iCLIP methods for nucleotide-resolution studies of protein-RNA interactions | Genome Biology | Full Text

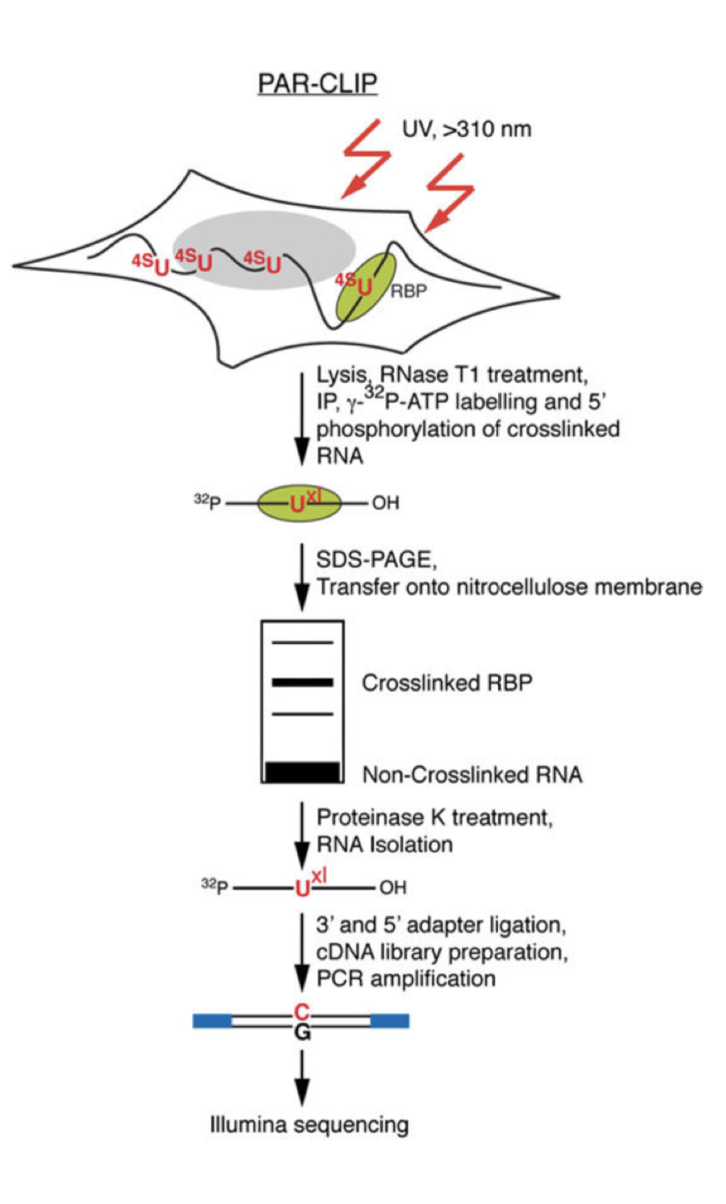
Transcriptome-wide Identification of RNA-Binding Protein and MicroRNA Target Sites by PAR-CLIP: Cell

Pipeline for extracting MIWI CLIP-Seq features (see Section 2). The RNA... | Download Scientific Diagram

omniCLIP: probabilistic identification of protein-RNA interactions from CLIP-seq data | Genome Biology | Full Text

PAR-CLIP and Streamlined Small RNA cDNA Library Preparation Protocol for the Identification of RNA Binding Protein Target Sites | RNA-Seq Blog




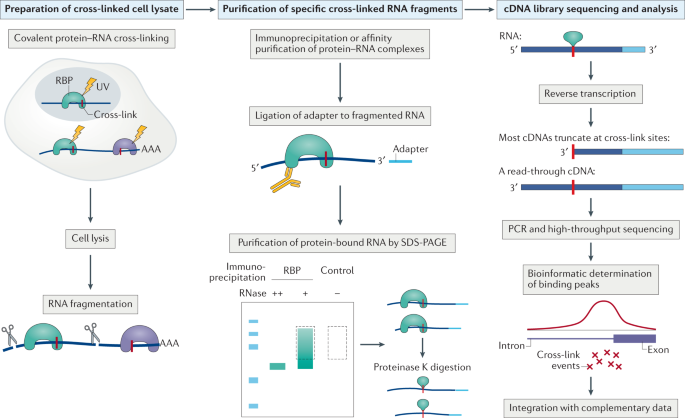
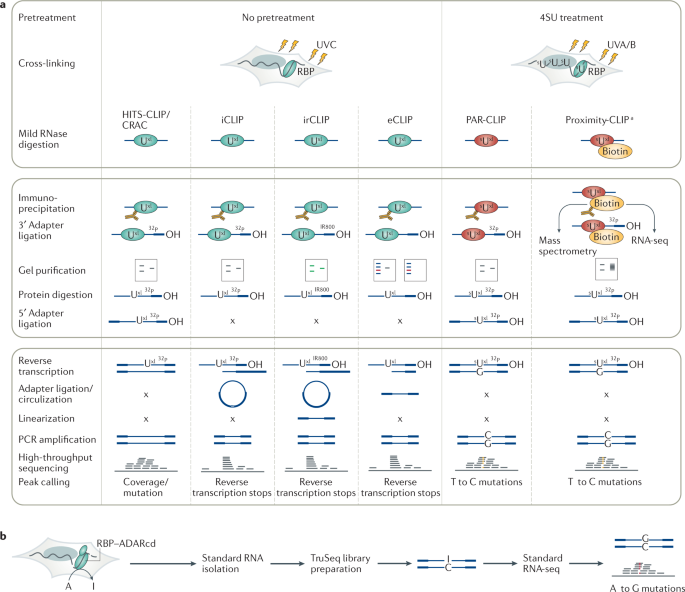
![PDF] The Future of Cross-Linking and Immunoprecipitation (CLIP). | Semantic Scholar PDF] The Future of Cross-Linking and Immunoprecipitation (CLIP). | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/f5ac369792eba3213c7d855f6e121dfcd9c4be6e/3-Figure1-1.png)
-2.jpg)